Doing CO1 and p47 PCRs on DNA from Single Fly DNA Extractions and extractions to Test for DiNV Infection
Samples for PCR in the layout they were in for the plates:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | 119 | 56 | 30 | 40 | 63 | 59 | 35 | 38 | 92 | 64 | 3 | 102 |
| B | 98 | 106 | 117 | 81 | 11 | 100 | 77 | 68 | 57 | 79 | 9 | 34 |
| C | 28 | 13 | 8 | 39 | 37 | 18 | 89 | 65 | 75 | 76 | 2 | 54 |
| D | 25 | 15 | 17 | 113 | 91 | 74 | 53 | 103 | 22 | 66 | 111 | 27 |
| E | 52 | 115 | 41 | 104 | 1 | 86 | 58 | 70 | 120 | 48 | 7 | 101 |
| F | 4 | 29 | 94 | 71 | 36 | 32 | 110 | 33 | 107 | 118 | 16 | 96 |
| G | 31 | 108 | 44 | 21 | 49 | 26 | 67 | 88 | 55 | |||
| H |
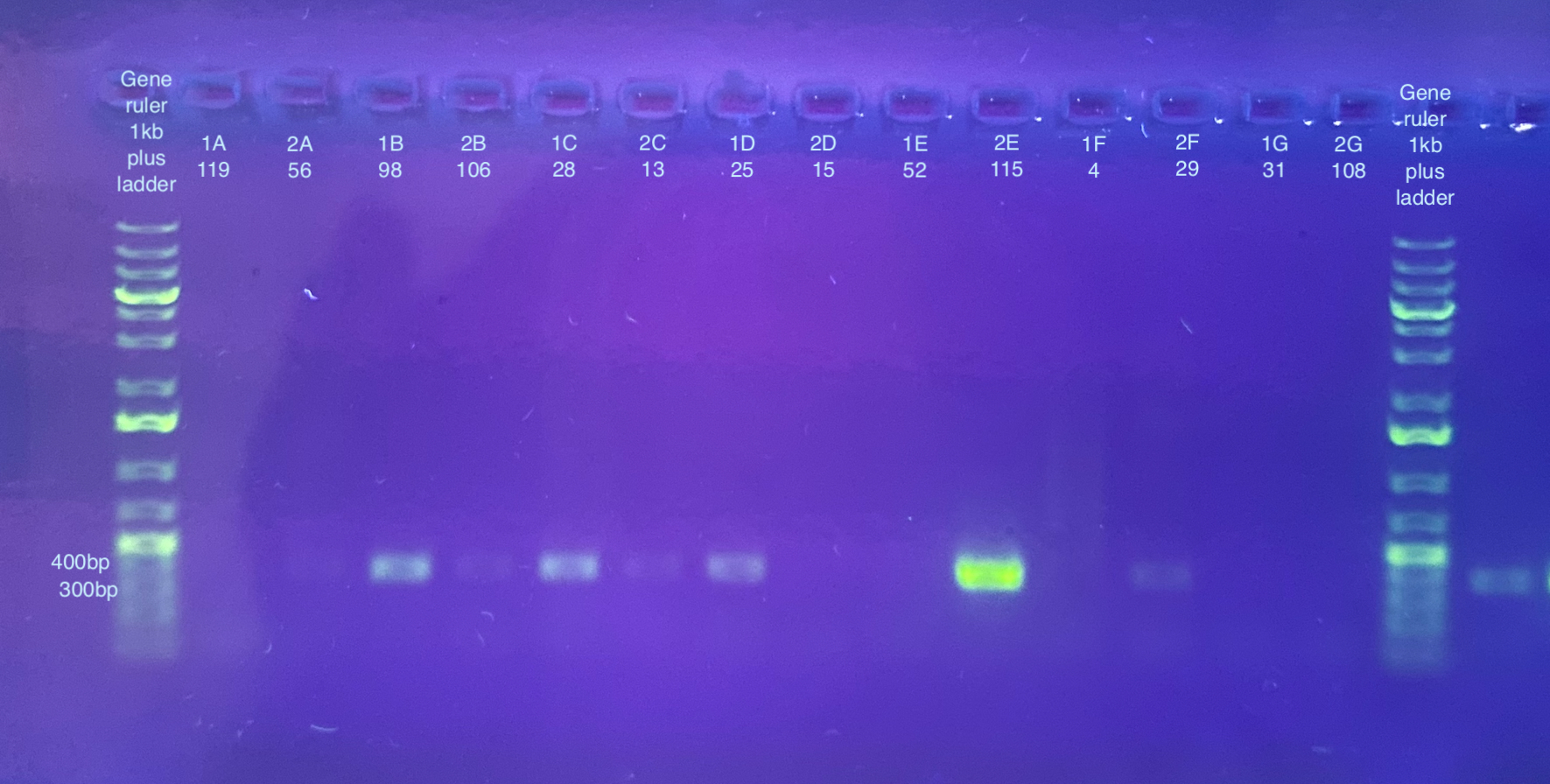
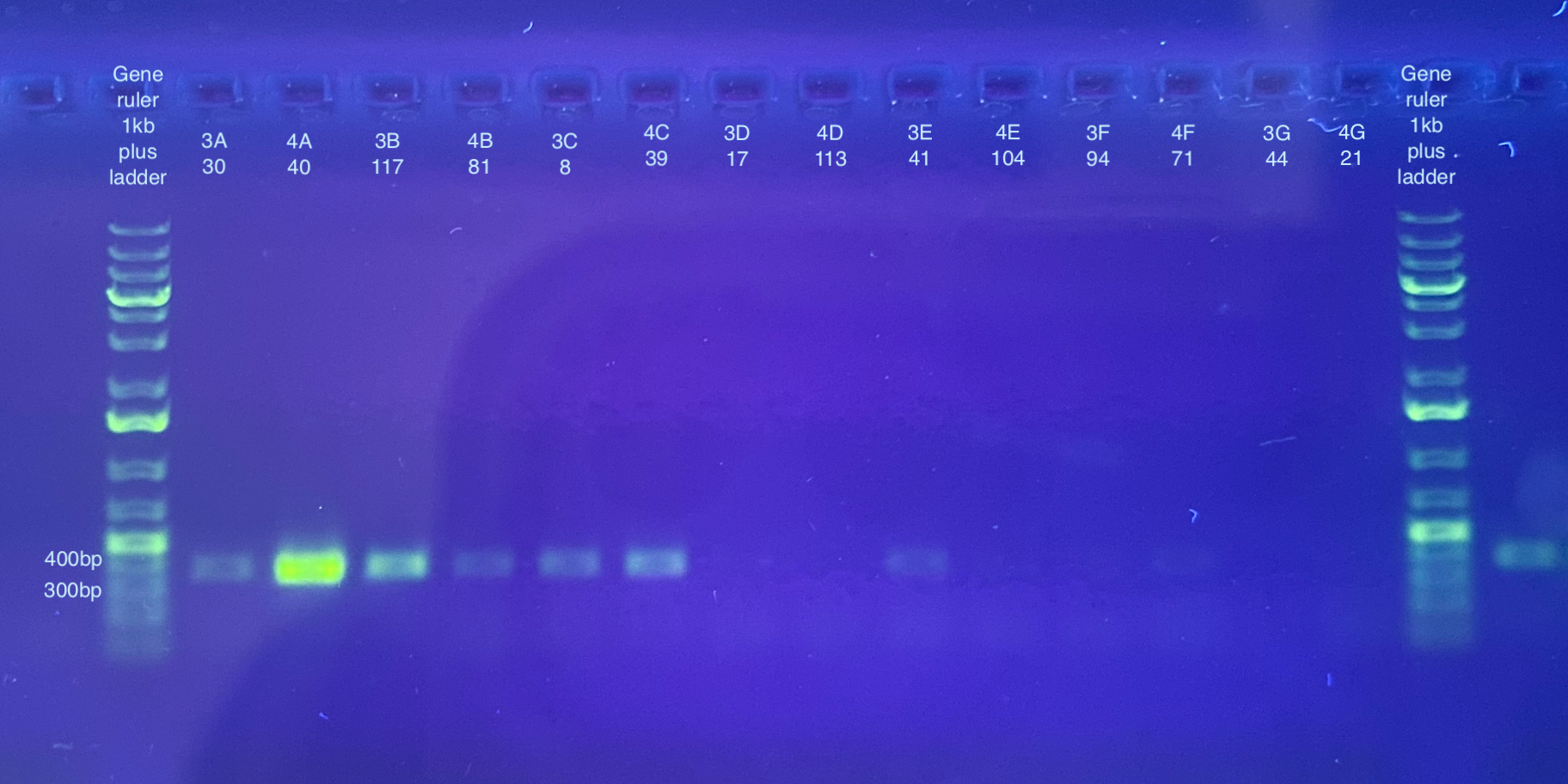
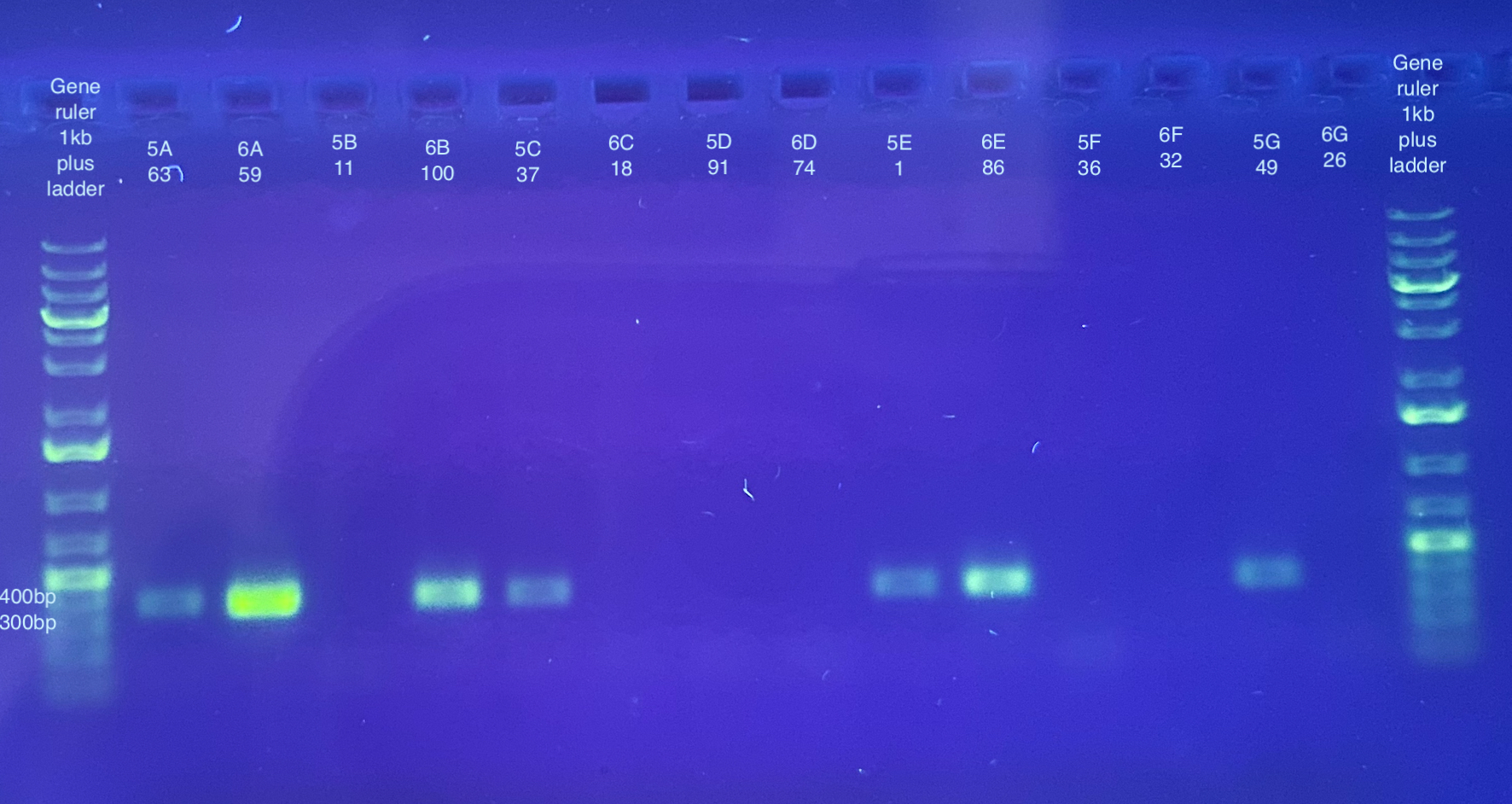
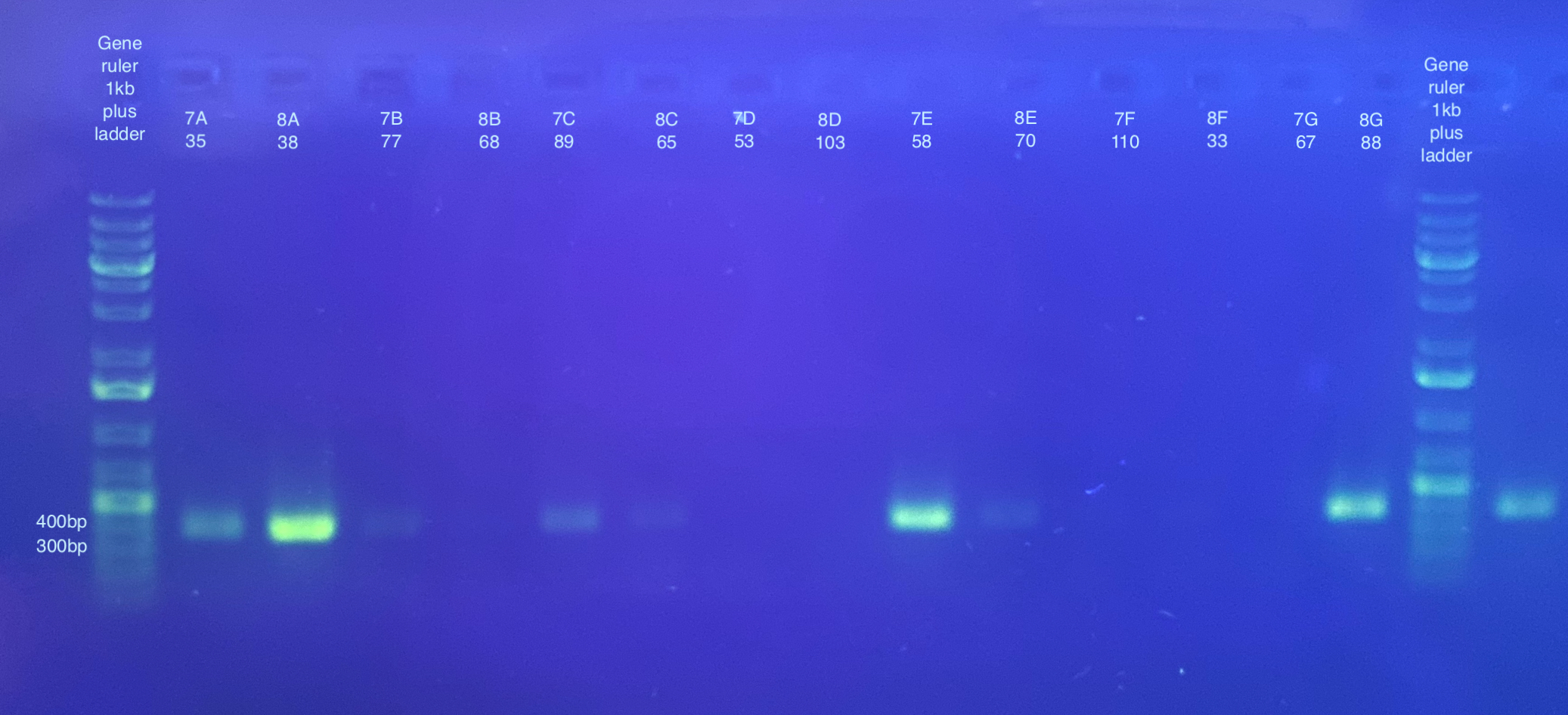
PCR followed general PCR protocol using the CO1 and p47 primers, and n number of 90, and cycling conditions indicated here. Note that I forgot to do a positive and negative control for these. I also became concerned that there was some splashing between samples when vortexing the plate because the seal may have not been fully on. I will likely have to do at least the p47 PCRs again.
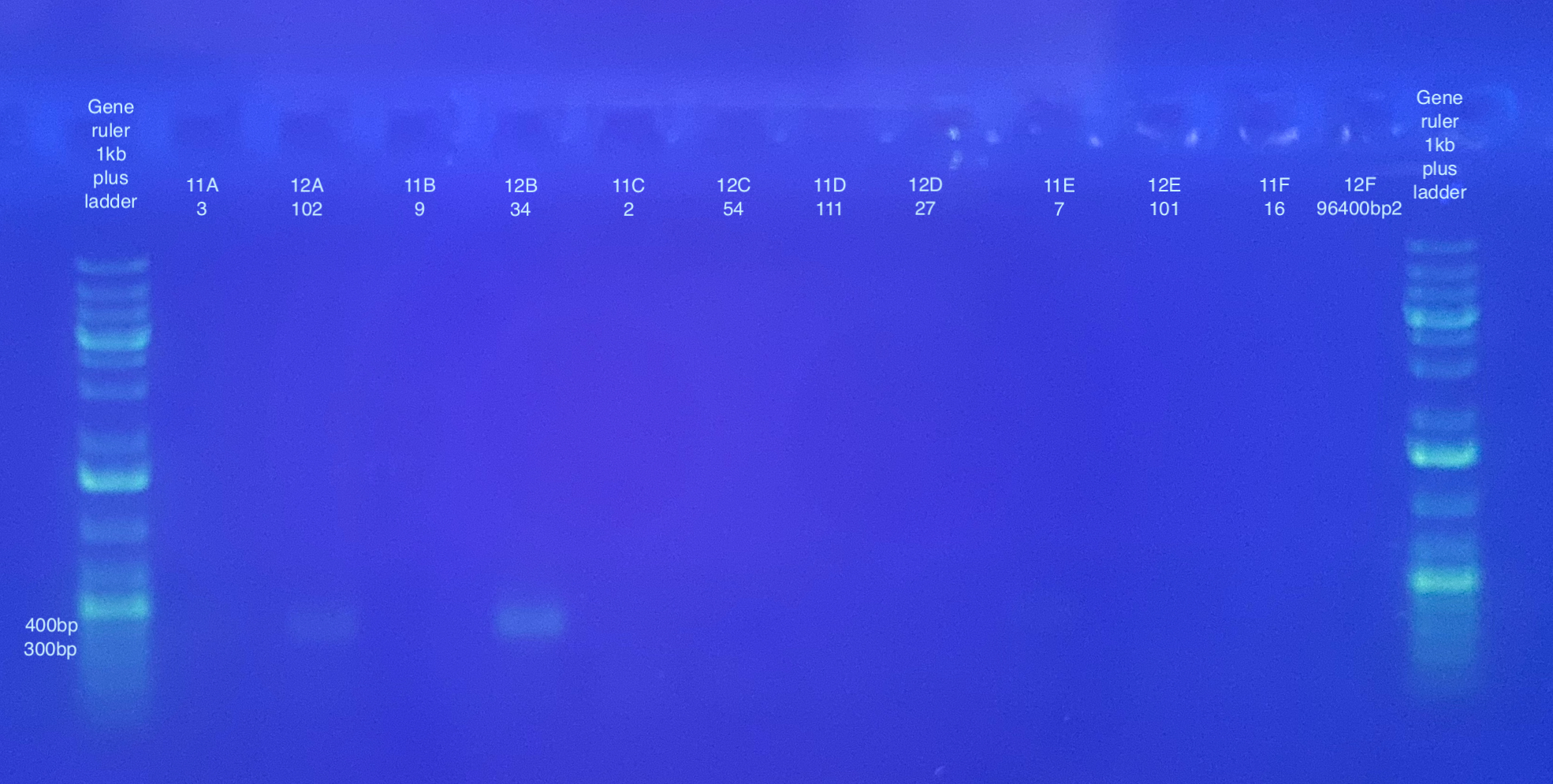
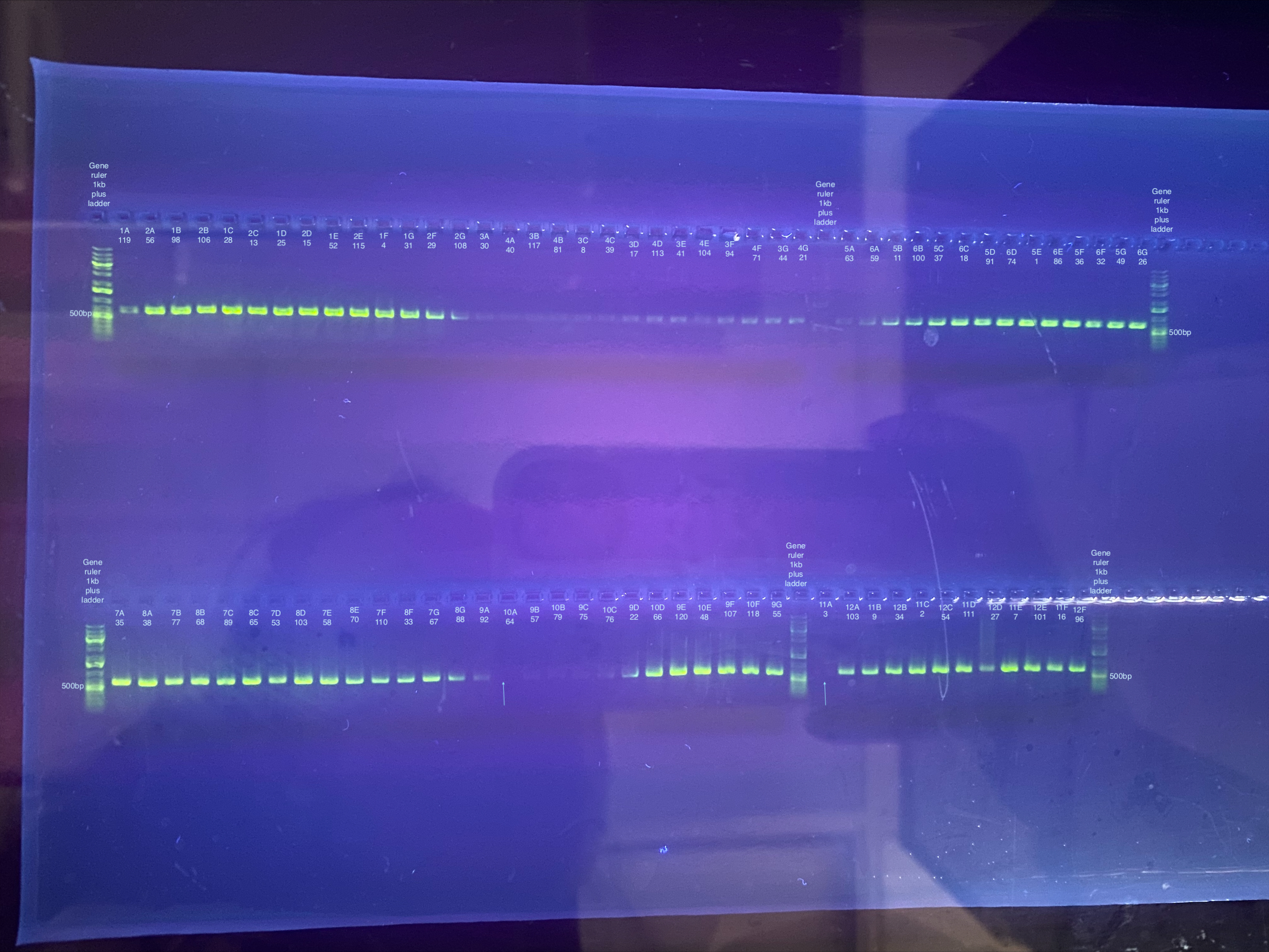
Separate gels run for both primer sets, 1% gel, 90V, 30 minutes for CO1, 40 minutes for p47. Note that the stain did not mix well for the CO1 gel, but that basically all samples seemed to have a band anyways.
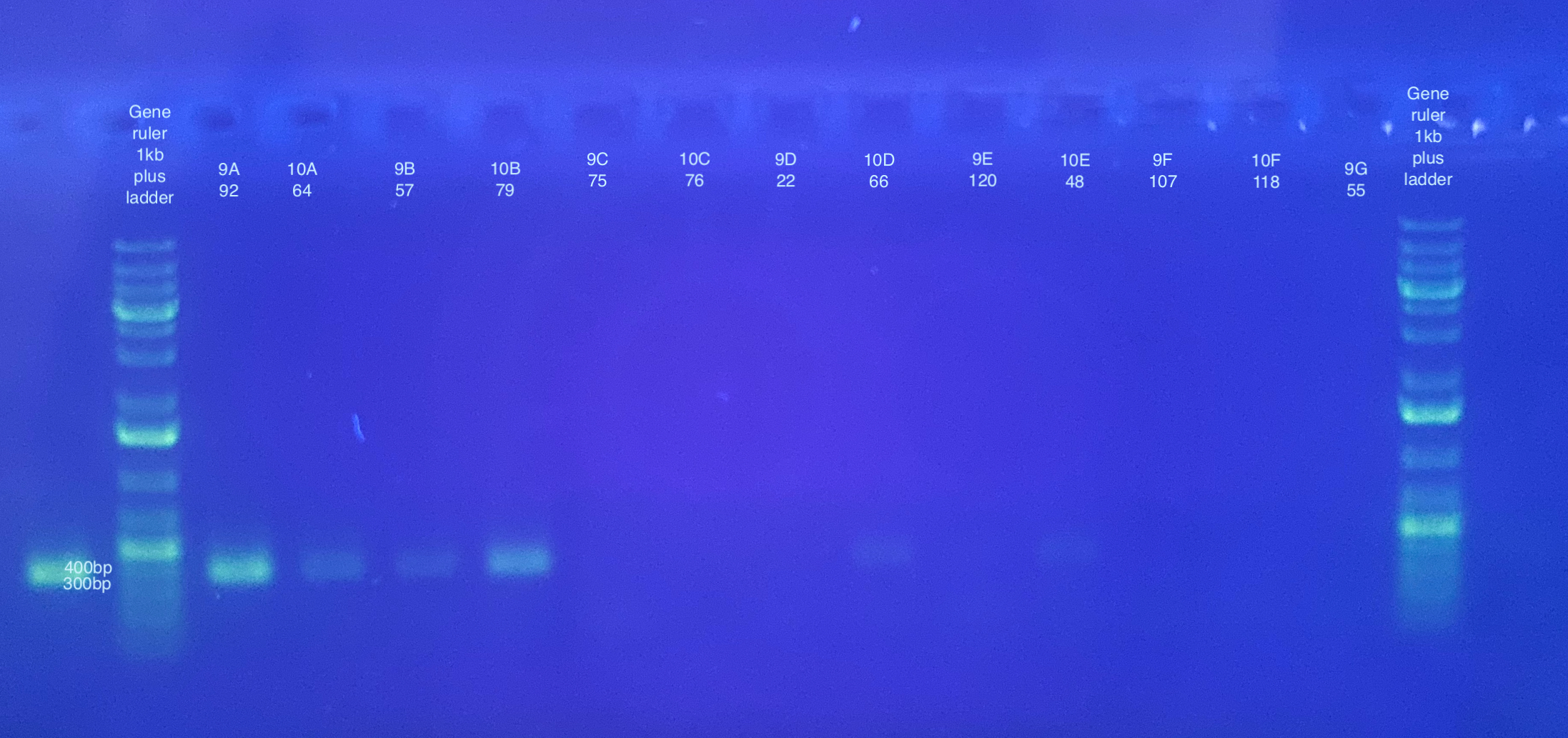
CO1:

p47 gel split up into multiple pictures: